|
| | hivpopulation (int N=0, int rng_seed=0, double mutation_rate=3e-5, double coinfection_rate=1e-2, double crossover_rate=1e-3) |
| | Construct a HIV population with certain parameters. More...
|
| |
| virtual | ~hivpopulation () |
| | Destructor. More...
|
| |
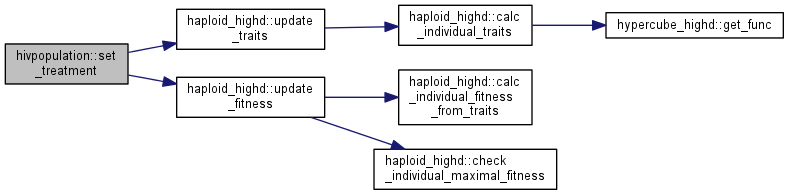
| void | set_treatment (double t) |
| |
| double | get_treatment () |
| |
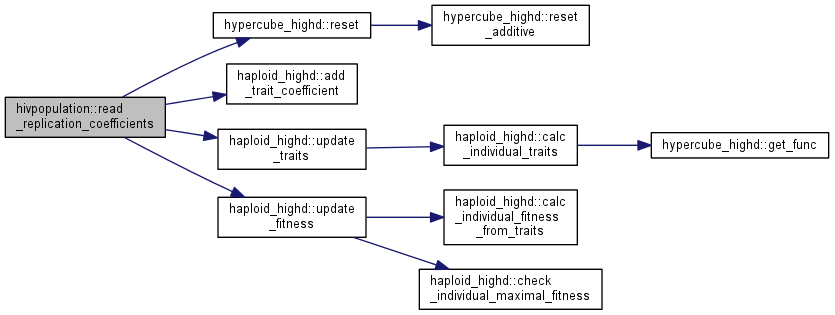
| int | read_replication_coefficients (istream &model) |
| |
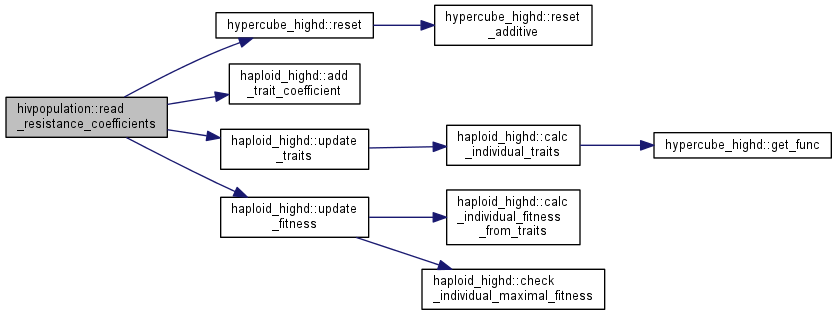
| int | read_resistance_coefficients (istream &model) |
| |
| int | write_genotypes (ostream &out_genotypes, int sample_size, string gt_label="", int start=0, int length=0) |
| |
| | haploid_highd (int L=0, int rng_seed=0, int number_of_traits=1, bool all_polymorphic=false) |
| | Default constructor. More...
|
| |
| virtual | ~haploid_highd () |
| | Destructor. More...
|
| |
| double | get_mutation_rate () |
| |
| void | set_mutation_rate (double m) |
| |
| bool | is_all_polymorphic () |
| |
| vector< poly_t > | get_polymorphisms () |
| |
| vector< poly_t > | get_fixed_mutations () |
| |
| vector< int > | get_number_of_mutations () |
| |
| int | L () |
| |
| int | get_number_of_loci () |
| |
| int | N () |
| |
| int | get_population_size () |
| |
| int | get_generation () |
| |
| void | set_generation (int g) |
| |
| int | get_number_of_clones () |
| |
| int | get_number_of_traits () |
| |
| double | get_participation_ratio () |
| |
| int | set_allele_frequencies (double *frequencies, unsigned long N) |
| | Initialize population in linkage equilibrium. More...
|
| |
| int | set_genotypes (vector< genotype_value_pair_t > gt) |
| | Initialize the population with genotype counts. More...
|
| |
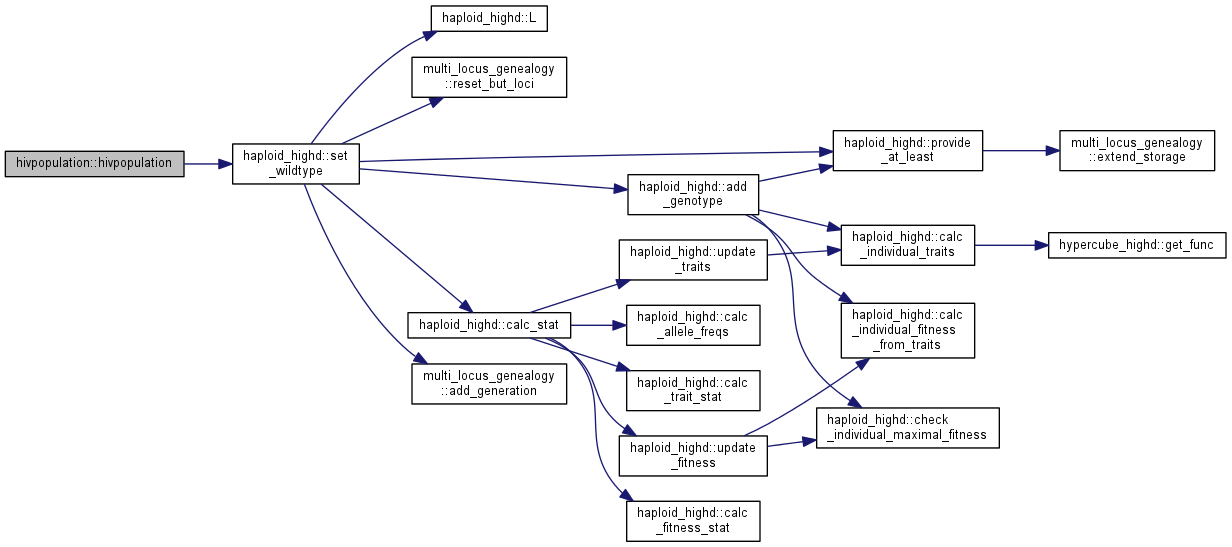
| int | set_wildtype (unsigned long N) |
| | Initialize a wildtype population (00...0) More...
|
| |
| int | track_locus_genealogy (vector< int > loci) |
| | Designates as set of loci to have their genealogy tracked. More...
|
| |
| void | add_genotype (boost::dynamic_bitset<> genotype, int n=1) |
| | Add the genotype specified by a bitset to the current population in in n copies. More...
|
| |
| int | add_trait_coefficient (double value, vector< int > loci, int t=0) |
| |
| void | clear_trait (int t=0) |
| |
| void | clear_traits () |
| |
| void | set_random_trait_epistasis (double epistasis_std, int traitnumber=0) |
| |
| int | add_fitness_coefficient (double value, vector< int > loci) |
| |
| void | clear_fitness () |
| |
| void | set_random_epistasis (double epistasis_std) |
| |
| int | evolve (int gen=1) |
| | Evolve for some generations under the specified conditions. More...
|
| |
| int | bottleneck (int size_of_bottleneck) |
| | Cause a bottleneck in the population size. More...
|
| |
| unsigned int | flip_single_locus (int locus) |
| | Flip a spin at a specific locus in random individual. More...
|
| |
| void | calc_stat () |
| | Calculate trait and fitness statistics and allele frequences. More...
|
| |
| void | unique_clones () |
| | Remove duplicate clones. More...
|
| |
| vector< int > | get_nonempty_clones () |
| | Obtain a list of the good clones. More...
|
| |
| int | random_clone () |
| | Get a random clone from the population. More...
|
| |
| int | random_clones (unsigned int n_o_individuals, vector< int > *sample) |
| | Sample random individuals from the population. More...
|
| |
| string | get_genotype_string (unsigned int i) |
| |
| int | distance_Hamming (unsigned int clone1, unsigned int clone2, vector< unsigned int * > *chunks=NULL, unsigned int every=1) |
| |
| int | distance_Hamming (boost::dynamic_bitset<> gt1, boost::dynamic_bitset<> gt2, vector< unsigned int * > *chunks=NULL, unsigned int every=1) |
| | Calculate Hamming distance between two sequences. More...
|
| |
| stat_t | get_diversity_statistics (unsigned int n_sample=1000) |
| | Calculate diversity in the current population (Hamming distance between pairs of sequences) More...
|
| |
| stat_t | get_divergence_statistics (unsigned int n_sample=1000) |
| | Calculate mean and variance of the divergence from the [00...0] bitset. More...
|
| |
| double | get_allele_frequency (int l) |
| |
| double | get_derived_allele_frequency (int l) |
| |
| double | get_pair_frequency (int locus1, int locus2) |
| | Get the joint frequency of two alleles. More...
|
| |
| vector< double > | get_pair_frequencies (vector< vector< int > > *loci) |
| | Get the joint frequency of two alleles, for a vector of allele pairs. More...
|
| |
| double | get_chi (int l) |
| |
| double | get_derived_chi (int l) |
| |
| double | get_chi2 (int locus1, int locus2) |
| |
| double | get_LD (int locus1, int locus2) |
| |
| double | get_moment (int locus1, int locus2) |
| |
| void | set_trait_weights (double *weights) |
| |
| double | get_trait_weight (int t) |
| |
| double | get_fitness (int n) |
| |
| int | get_clone_size (int n) |
| |
| double | get_trait (int n, int t=0) |
| |
| vector< coeff_t > | get_trait_epistasis (int t=0) |
| |
| stat_t | get_fitness_statistics () |
| |
| stat_t | get_trait_statistics (int t=0) |
| |
| double | get_trait_covariance (int t1, int t2) |
| |
| double | get_max_fitness () |
| |
| void | update_traits () |
| | For each clone, recalculate its traits. More...
|
| |
| void | update_fitness () |
| | For each clone, update fitness assuming traits are already up to date. More...
|
| |
| int | get_divergence_histogram (gsl_histogram **hist, unsigned int bins=10, vector< unsigned int * > *chunks=NULL, unsigned int every=1, unsigned int n_sample=1000) |
| | Get histogram of divergence from the [00...0] bitset. More...
|
| |
| int | get_diversity_histogram (gsl_histogram **hist, unsigned int bins=10, vector< unsigned int * > *chunks=NULL, unsigned int every=1, unsigned int n_sample=1000) |
| | Get histogram of diversity in the population (mutual Hamming distance) More...
|
| |
| int | get_fitness_histogram (gsl_histogram **hist, unsigned int bins=10, unsigned int n_sample=1000) |
| | Calculate histogram of fitness from traits. More...
|
| |
| int | print_allele_frequencies (ostream &out) |
| | Print all allele frequencies into a stream provided. More...
|
| |
| int | read_ms_sample (istream >s, int skip_locus, int multiplicity) |
| | Read the output of Hudson's ms and use it to initialize the genotype distribution. More...
|
| |
| int | read_ms_sample_sparse (istream >s, int skip_locus, int multiplicity, int distance) |
| | Read the output of Hudson's ms and use it to initialize the genotype distribution. More...
|
| |
|
| virtual void | calc_individual_fitness_from_traits (clone_t *tempgt) |
| | Set the fitness from replication and resistance for a single clone. More...
|
| |
| int | get_random_seed () |
| | Get a random seed from /dev/urandom. More...
|
| |
| void | produce_random_sample (int size=1000) |
| | Produce and store a random sample of the population for stochastic processes. More...
|
| |
| int | mutate () |
| | Mutate random clones at random loci (all loci) More...
|
| |
| int | select_gametes () |
| | Generate offspring according to fitness (selection) and segregate some for sexual mating. More...
|
| |
| double | relaxation_value () |
| | Get the log of the exp-average fitness plus relaxation term. More...
|
| |
| double | get_logmean_expfitness () |
| | Get the population exp-average of fitness, used for keeping the population size fixed. More...
|
| |
| unsigned int | flip_single_locus (unsigned int clonenum, int locus) |
| | Flip a spin (locus) in individual. More...
|
| |
| void | shuffle_genotypes () |
| |
| int | new_generation () |
| |
| int | partition_cumulative (vector< unsigned int > &partition_cum) |
| | Calculate the cumulative partition of sequences into clones. More...
|
| |
| int | provide_at_least (int n) |
| | : allocates memory for a sufficient number of clones More...
|
| |
| void | calc_allele_freqs () |
| | calculate and store allele frequencies More...
|
| |
| void | reassortment_pattern () |
| | Produce a random reassortement pattern. More...
|
| |
| void | crossover_pattern () |
| | Choose a number of crossover points and produce a crossover pattern. More...
|
| |
| int | add_recombinants () |
| | Pair and mate sexual gametes. More...
|
| |
| int | recombine (int parent1, int parent2) |
| | Recombine two genotypes parent1 and parent2 to produce two new genotypes. More...
|
| |
| int | recombine_crossover (int parent1, int parent2, int ng) |
| |
| void | calc_fitness_stat () |
| | Calculate and store fitness population statistics. More...
|
| |
| void | calc_trait_stat () |
| | Calculate and store trait population statistics and covariances. More...
|
| |
| void | calc_individual_traits (clone_t &tempgt) |
| | Calculate traits of the chosen clone. More...
|
| |
| void | calc_individual_fitness (clone_t &tempgt) |
| | Calculate fitness of a particular clone. More...
|
| |
| void | calc_individual_traits (int clonenum) |
| |
| void | calc_individual_fitness (int clonenum) |
| |
| void | check_individual_maximal_fitness (clone_t &tempgt) |
| |
| double | get_trait_difference (clone_t &tempgt1, clone_t &tempgt2, vector< int > &diffpos, int traitnum) |
| | Calculate trait difference between two clones. More...
|
| |
| virtual void | calc_individual_fitness_from_traits (clone_t &tempgt) |
| | Calculate fitness from traits of the chosen clone. More...
|
| |
| virtual void | calc_individual_fitness_from_traits (int clonenum) |
| |
| void | add_clone_to_genealogy (int locus, int dest, int parent, int left, int right, int cs, int n) |
| |
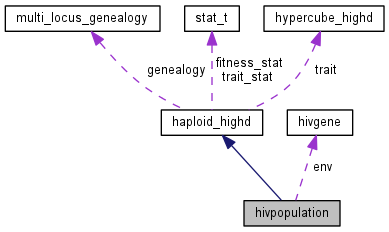
HIV population with facultative drug treatment.
This class exemplifies the haploid_highd base class. It mainly adds one trait, "treatment", which is the same for all individuals and represents the presence or absence of drug treatment (in a continuous manner, \(0 \leq \) treatment \(\leq 1\)).
The replication capacity in absence of drug is encoded in the first trait. The drug resistance phenotype is represented by the second trait. Fitness is computed from traits as follows:
f[trait] = trait[0] + treatment * trait[1]
Moreover, this class fixes the length of the genome to exactly 10000 sites.
 Public Member Functions inherited from haploid_highd
Public Member Functions inherited from haploid_highd Public Attributes inherited from haploid_highd
Public Attributes inherited from haploid_highd Protected Member Functions inherited from haploid_highd
Protected Member Functions inherited from haploid_highd Protected Attributes inherited from haploid_highd
Protected Attributes inherited from haploid_highd