|
FFPopSim
2.0
Library for Population Genetics
|
|
FFPopSim
2.0
Library for Population Genetics
|
Low-dimensional population evolving on the hypercube_lowd. More...
#include <ffpopsim_lowd.h>
Public Member Functions | |
| haploid_lowd (int L=1, int rng_seed=0) | |
| Default constructor. More... | |
| virtual | ~haploid_lowd () |
| Default destructor. More... | |
| int | L () |
| int | get_number_of_loci () |
| double | N () |
| double | get_population_size () |
| double | get_generation () |
| void | set_generation (double g) |
| double | get_mutation_rate (int locus, int direction) |
| int | get_recombination_model () |
| double | get_recombination_rate (int locus) |
| Get the recombination rate between a locus and the next one. More... | |
| int | set_allele_frequencies (double *frequencies, unsigned long N) |
| Initialize population in linkage equilibrium. More... | |
| int | set_genotypes (vector< index_value_pair_t > gt) |
| Initialize the population with genotype counts. More... | |
| int | set_wildtype (unsigned long N) |
| Initialize a wildtype population (00...0) More... | |
| int | set_recombination_model (int rec_model) |
| Set a new recombination model. More... | |
| int | set_recombination_rates (double *rec_rates, int rec_model=-1) |
| calculate recombination patterns More... | |
| int | set_mutation_rates (double m) |
| Set a uniform mutation rate for all loci and both directions. More... | |
| int | set_mutation_rates (double m1, double m2) |
| Set two mutation rates (forward / backward) for all loci. More... | |
| int | set_mutation_rates (double *m) |
| Set mutation rates (locus specific, both directions the same) More... | |
| int | set_mutation_rates (double **m) |
| Set mutation rates (locus and direction specific) More... | |
| int | evolve (int gen=1) |
| Evolve the population for some generations. More... | |
| int | evolve_norec (int gen=1) |
| Evolve the population for some generations, without recombination. More... | |
| int | evolve_deterministic (int gen=1) |
| Evolve the population for some generations, without resampling (deterministic) More... | |
| double | get_genotype_frequency (int genotype) |
| double | get_allele_frequency (int locus) |
| double | get_pair_frequency (int locus1, int locus2) |
| double | get_chi (int locus) |
| double | get_chi2 (int locus1, int locus2) |
| double | get_LD (int locus1, int locus2) |
| double | get_moment (int locus1, int locus2) |
| double | genotype_entropy () |
| Get the genotype entropy. More... | |
| double | allele_entropy () |
| Get the allele entropy. More... | |
| double | get_fitness (int genotype) |
| stat_t | get_fitness_statistics () |
| Get fitness mean and variance in the population. More... | |
Public Attributes | |
| hypercube_lowd | fitness |
| hypercube_lowd | population |
| double | carrying_capacity |
| double | outcrossing_rate |
| bool | circular |
Protected Member Functions | |
| int | get_random_seed () |
| Get a random seed from /dev/urandom. More... | |
| int | select () |
| Selection step. More... | |
| int | mutate () |
| Mutation step. More... | |
| int | recombine () |
| Recombination step. More... | |
| int | resample () |
| Resample the population according to the carrying capacity. More... | |
| int | set_recombination_rates_general (double *rec_rates) |
| Set the recombination rates for the general model. More... | |
| int | set_recombination_patterns (vector< index_value_pair_t > iv) |
| set recombination patterns with a list of index value pairs More... | |
| int | marginalize_recombination_patterns () |
| Starting from a fully specified set of probabilities of recombination patterns, compute all sub patterns. More... | |
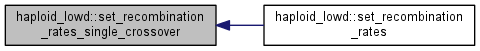
| int | set_recombination_rates_single_crossover (double *rec_rates) |
| Set the recombination rates for the single crossover model. More... | |
| int | calculate_recombinants_free () |
| Calculate the recombinant distribution for the free recombination case. More... | |
| int | calculate_recombinants_single () |
| Calculate the recombinant distribution for the single crossover case. More... | |
| int | calculate_recombinants_general () |
| Calculate the recombinant distribution for the general case. More... | |
Protected Attributes | |
| gsl_rng * | rng |
| int | seed |
| hypercube_lowd | recombinants |
| hypercube_lowd | mutants |
| double ** | recombination_patterns |
| int | recombination_model |
| int | number_of_loci |
| double | population_size |
| int | generation |
| double | long_time_generation |
| double ** | mutation_rates |
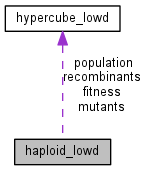
Low-dimensional population evolving on the hypercube_lowd.
This class enables simulation of short genomes ( \(L \lesssim 20\)) but potentially large populations. Random mutation, recombination and selection are supported. A number of properties of the population can be obtained using methods of this class, including:
| haploid_lowd::haploid_lowd | ( | int | L_in = 1, |
| int | rng_seed = 0 |
||
| ) |
Default constructor.
| L_in | number of loci (at least 1) |
| rngseed | seed for the random number generator. If this is zero, time(NULL)+getpid() is used. |

|
virtual |
Default destructor.
Release memory.
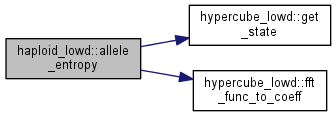
| double haploid_lowd::allele_entropy | ( | ) |
Get the allele entropy.
The allele entropy is defined as follows:
\[ S := - \sum_{i=1}^L \nu_i\log \nu_i + (1 - \nu_i)\log (1 - \nu_i), \]
where \(\nu_i\) is the frequency of the i-th allele.
|
protected |
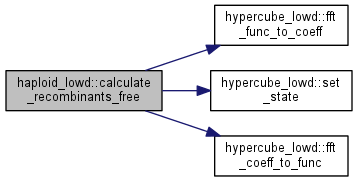
Calculate the recombinant distribution for the free recombination case.
Almost the same as for the more general case below, but kept separate for performance reasons - this is the most expensive part (3^L).
|
protected |
Calculate the recombinant distribution for the general case.
Calculate the distribution after recombination assumed in random mating with pairs sampled with replacement.
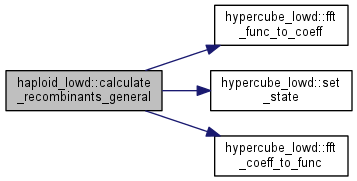
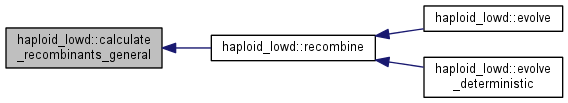
|
protected |
Calculate the recombinant distribution for the single crossover case.
Calculate the distribution after recombination assumed in random mating with pairs sampled with replacement.
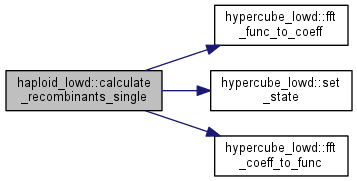
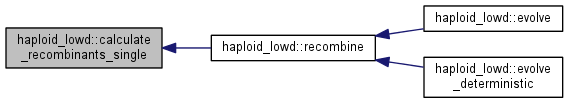
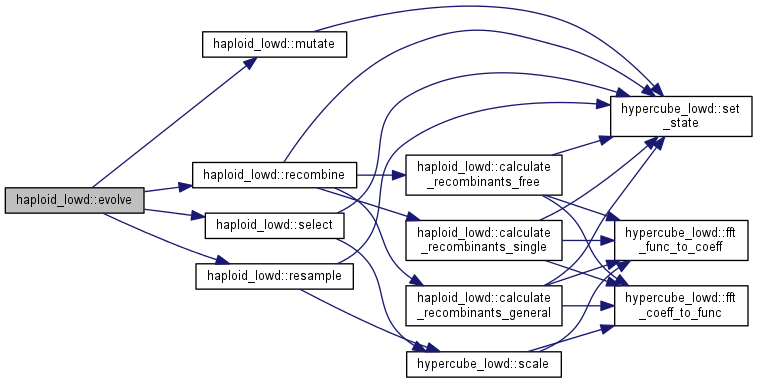
| int haploid_lowd::evolve | ( | int | gen = 1 | ) |
Evolve the population for some generations.
| gen | number of generations |
Note: the order of selection, mutation, recombination, and resampling could be changed according to needs and beliefs. Note that only recombination calculates the inverse fourier transform of the population. It does so BEFORE the recombination step. To evaluate all allele frequencies and linkage disequilibria, call population.fft_func_to_coeff()
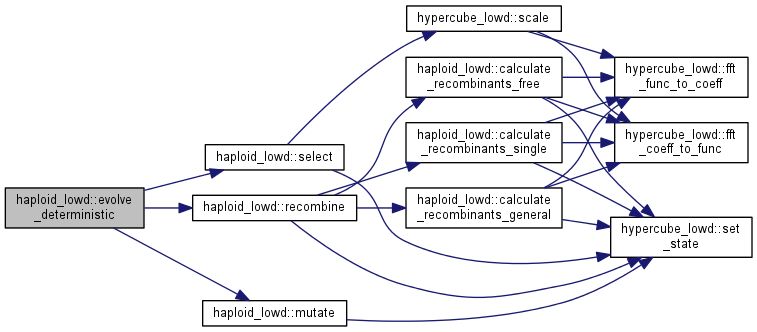
| int haploid_lowd::evolve_deterministic | ( | int | gen = 1 | ) |
Evolve the population for some generations, without resampling (deterministic)
| gen | number of generations |
Note: the order of selection, mutation, and recombination could be changed according to needs and beliefs. Note that only recombination calculates the inverse fourier transform of the population. It does so BEFORE the recombination step. To evaluate all allele frequencies and linkage disequilibria, call population.fft_func_to_coeff()
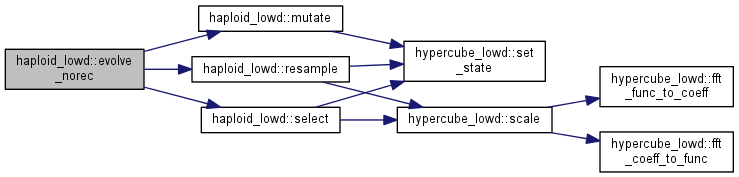
| int haploid_lowd::evolve_norec | ( | int | gen = 1 | ) |
Evolve the population for some generations, without recombination.
| gen | number of generations |
Note: the order of selection, mutation, and resampling could be changed according to needs and beliefs.
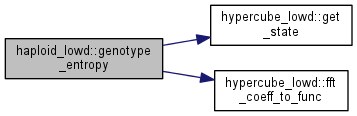
| double haploid_lowd::genotype_entropy | ( | ) |
Get the genotype entropy.
The genotype entropy is defined as follows:
\[ S := - \sum_{g} \nu_g \cdot \log \nu_g, \]
where \(g\) runs over all possible genomes, and \(\nu_g\) is the frequency of that genome in the population.
|
inline |
|
inline |
|
inline |
|
inline |
| stat_t haploid_lowd::get_fitness_statistics | ( | ) |
Get fitness mean and variance in the population.

|
inline |
|
inline |
|
inline |
|
inline |
|
inline |
|
inline |
|
inline |
|
inline |
|
protected |
Get a random seed from /dev/urandom.

|
inline |
| double haploid_lowd::get_recombination_rate | ( | int | locus | ) |
Get the recombination rate between a locus and the next one.
| locus | the locus AFTER which the recombination rate is seeked |
Note: for SINGLE_CROSSOVER, the recombination rate after the last locus is the rate of no recombination at all.
|
inline |
|
protected |
Starting from a fully specified set of probabilities of recombination patterns, compute all sub patterns.
| @returns | zero if successful, error codes otherwise |

|
protected |
Mutation step.
Calculate the distribution of mutants and update the population distribution
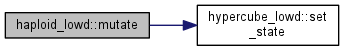
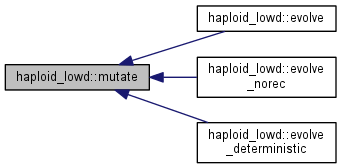
|
inline |
|
protected |
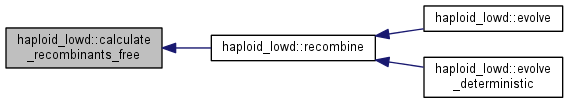
Recombination step.
Calculate the distribution of recombinants and update the population, a fraction (outcrossing_rate) of the population is replaced by the recombinant distribution, calculated differently depending on whether free recombination or general recombination is used
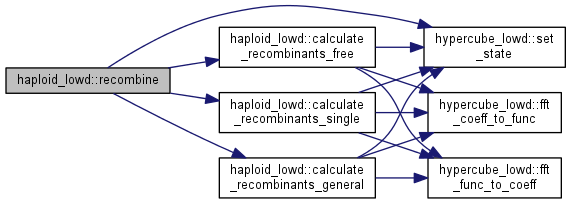
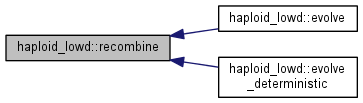
|
protected |
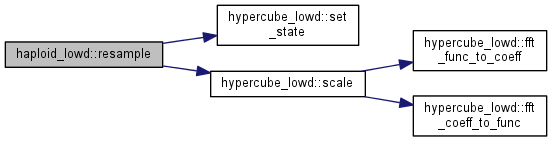
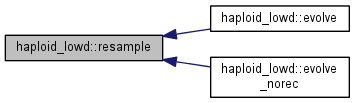
Resample the population according to the carrying capacity.
Note: genotypes with few individuals are sampled using the Poisson distribution, allowing for strict zero; genotypes with many individuals are resampled using a Gaussian distribution, for performance reasons.
|
protected |
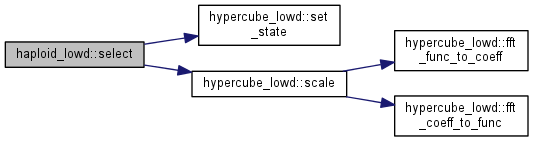
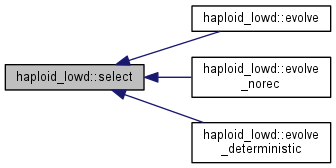
Selection step.
Note: Population distribution is reweighted with exp(fitness) and renormalized.
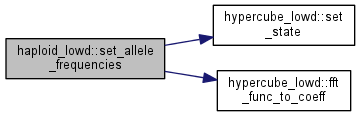
| int haploid_lowd::set_allele_frequencies | ( | double * | freq, |
| unsigned long | N_in | ||
| ) |
Initialize population in linkage equilibrium.
| freq | allele frequencies |
| N_in | population size |
Note: when this function is used to initialize the population, it is likely that the fitness distribution has a very large width. In turn, this can result in an immediate and dramatic drop in diversity within the first few generations. Please check fitness statistics before starting the evolution if this worries you.
Note: the population size will be set. If not set yet, the carrying capacity will be set to the same number.
|
inline |
| int haploid_lowd::set_genotypes | ( | vector< index_value_pair_t > | gt | ) |
Initialize the population with genotype counts.
| gt | vector of index_value_pair_t type with indices and counts |
Note: the population size will be set as the total sum of counts of all genotypes. If not set yet, the carrying capacity will be set to the same number.

| int haploid_lowd::set_mutation_rates | ( | double | m | ) |
Set a uniform mutation rate for all loci and both directions.
| m | mutation rate |
| int haploid_lowd::set_mutation_rates | ( | double | mforward, |
| double | mbackward | ||
| ) |
Set two mutation rates (forward / backward) for all loci.
| mforward | forward mutation rate |
| mbackward | backward mutation rate |
| int haploid_lowd::set_mutation_rates | ( | double * | m | ) |
Set mutation rates (locus specific, both directions the same)
| m | array of mutation rates |
| int haploid_lowd::set_mutation_rates | ( | double ** | m | ) |
Set mutation rates (locus and direction specific)
| m | array of mutation rates |
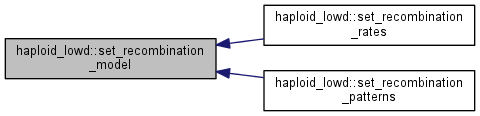
| int haploid_lowd::set_recombination_model | ( | int | rec_model | ) |
Set a new recombination model.
| rec_model | the new recombination model |
The allowed models are FREE_RECOMBINATION, SINGLE_CROSSOVER, and CROSSOVERS.
Note: this function call involves memory release and allocation.
|
protected |
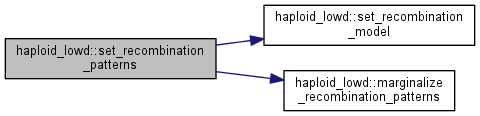
set recombination patterns with a list of index value pairs
| vector | of index_value_pair_t structures |
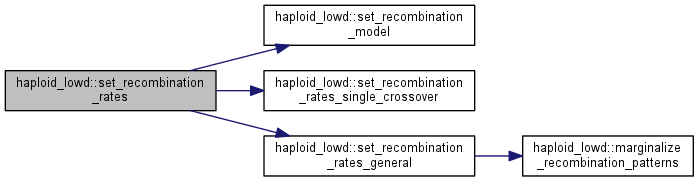
| int haploid_lowd::set_recombination_rates | ( | double * | rec_rates, |
| int | rec_model = -1 |
||
| ) |
calculate recombination patterns
| rec_rates | a vector of recombination rates. |
| rec_model | an int with the model of recombination to use |
Note: rec_rates must have length L-1 for linear chromosomes, length L for circular ones. Note: rec_model must be either CROSSOVERS or, for linear genomes, SINGLE_CROSSOVER
A routine the calculates the probability of all possible recombination patterns and subpatterns thereof from a vector of recombination rates (rec_rates) passed as argument. It allocated the memory ( \(3^L\) or \(L \: 2^L\)) and calculates the entire distribution.
Note: the default value for rec_model is the current recombination model or, if that is FREE_RECOMBINATION, then it's CROSSOVERS.
|
protected |
Set the recombination rates for the general model.
| rec_rates | array with the recombination rates |


|
protected |
Set the recombination rates for the single crossover model.
| rec_rates | array with the recombination rates |
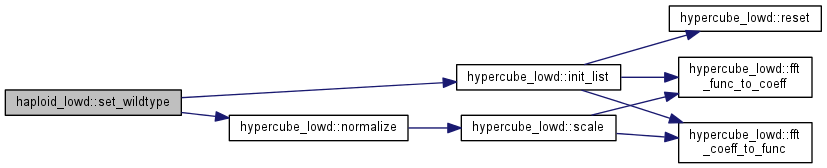
| int haploid_lowd::set_wildtype | ( | unsigned long | N_in | ) |
Initialize a wildtype population (00...0)
| N_in | number of individuals |
Note: the population size will be set. If not set yet, the carrying capacity will be set to the same number.
| double haploid_lowd::carrying_capacity |
| bool haploid_lowd::circular |
| hypercube_lowd haploid_lowd::fitness |
|
protected |
|
protected |
|
protected |
|
protected |
|
protected |
| double haploid_lowd::outcrossing_rate |
| hypercube_lowd haploid_lowd::population |
|
protected |
|
protected |
|
protected |
|
protected |
|
protected |
|
protected |